What life cycle traits can make a species more vulnerable to extinction?
by Judy Che-Castaldo on Mar 2, 2022(Blog post written by Haydée Hernández-Yáñez)
Anthropogenic threats to biodiversity are numerous and so are the number of species threatened by them. Conservation practitioners utilize the few resources available to them to create programs to protect species and where they live, but in many cases, it is difficult to decide where and when to allocate these resources. The field of conservation biology has worked extensively on the question that could aid in the decision making process: are there species more affected than others by anthropogenic threats? And how can we know which are these? It is certainly not a straight answer. Part of what complicates the matter is that species have intrinsic traits (life-history strategies, genetic) that allow them to respond differently to threats and therefore differ in their vulnerability to extinction. Studies have mostly concentrated in one to two taxa, probably because collecting species data is intensive, expensive, and may require long-term monitoring. A few other studies have circumvented this by using simulated data. The results of these studies linking intrinsic traits to vulnerability have been variable.
Taking advantage of the wealth of information contained in the global database that is COM(P)ADRE, my colleagues and I published a paper in PLOS ONE where we aimed to identify if some species are more vulnerable to extinction than others based on characteristics of their life cycle. Other studies have used lifespan, estimates of species abundance, survival rates and yet, to our knowledge, no systematic analyses with real demographic patterns and using the full species life cycle had been done. We gathered the demographic information from COM(P)ADRE from species within four different taxa: 46 tree species, 39 herbaceous perennials, 31 birds and 43 mammals, and used the IUCN Red List assessments as our proxy for vulnerability to extinction. We also obtained geolocation information for all species in the database and explored if most of the species are concentrated in biodiversity hotspots; and if a higher number of these species with an IUCN assessment are within or outside of hotspots.
Interestingly, we found there are demographic characteristics that do make certain taxa more vulnerable: herbaceous plants that reproduce later in life, birds that grow faster and produce more offspring, and mammals that are longer-lived (Fig. 1), tend to be more prone to extinction. These findings suggest that human disturbance does not affect all species equally thanks to their intrinsic biology. Subsequent studies should be done to understand why these patterns seem to be the most important predictors of extinction risk, as we only looked for the relationship between life-history traits and risk.
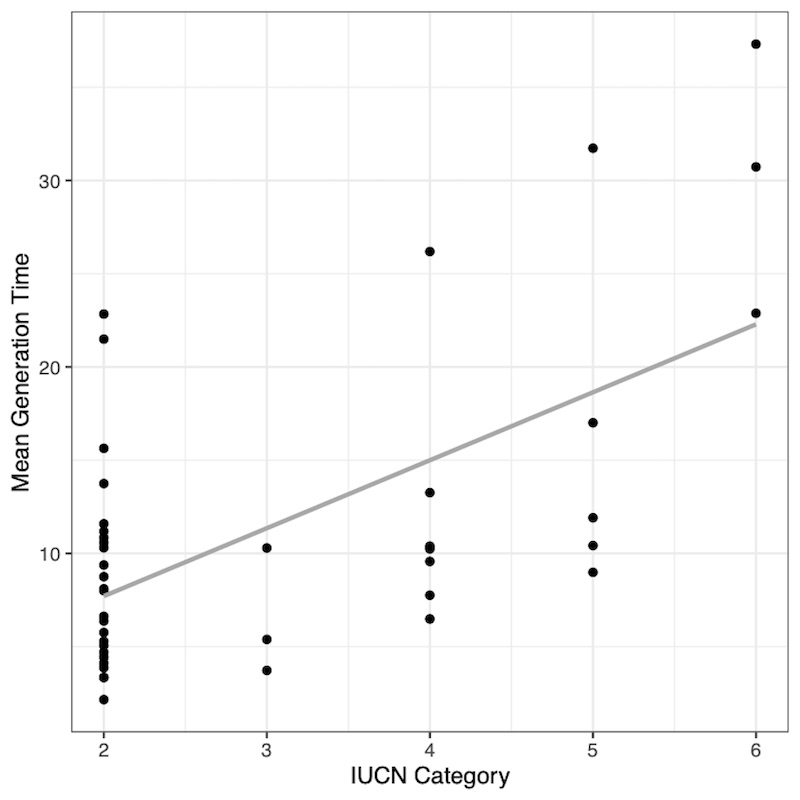
Figure 1. Mammals: the longer the generation time, the higher the IUCN category.
We also showed that COM(P)ADRE’s recorded species are not mainly concentrated in hotspots, yet most of the species in hotspots have an IUCN red list assessment (except for herbaceous perennials; Fig. 2).
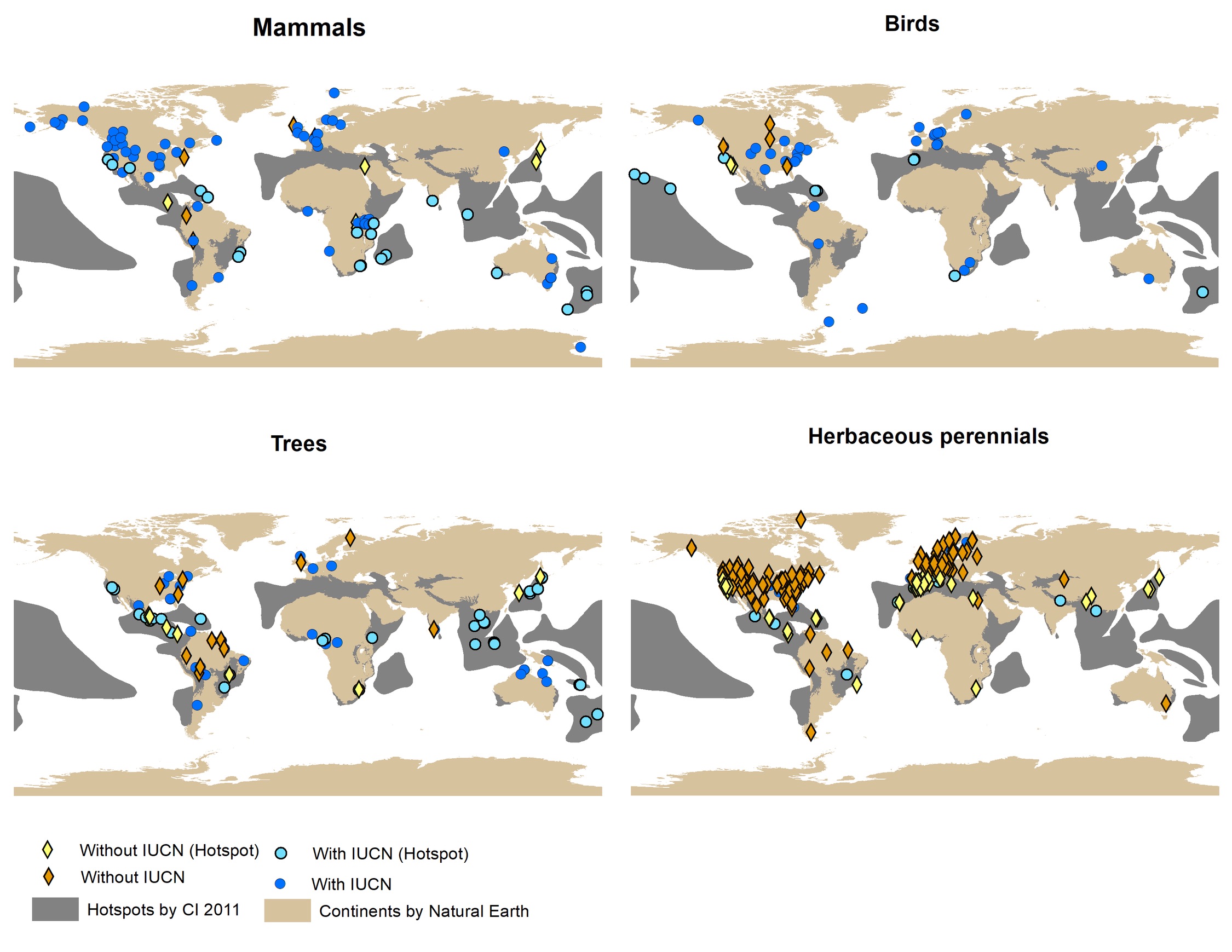
Figure 2. Spatial distribution of species in the COMPADRE and COMADRE matrix databases.
These results show that we can use demographic patterns to detect species vulnerability to extinction, especially when estimates of abundance and trends are often not available and costly to obtain. This information can be used to find species that may be prime candidates for extinction to concentrate restoration efforts on those species. Our study also demonstrates an added value for the COM(P)ADRE data: many species (especially herbaceous perennial plants) that do not have an IUCN Red List assessment already have demographic information in this database, and can benefit from it to determine their potential risk of extinction.


